Recombinant DNA Technology
Contents for Tables:-
- Introduction
- Definition to Recombinant DNA Technology
- Historical Context and Development
- Key Concepts in Recombinant DNA Technology
- Molecular Genetics Overview
- Fundamental of Technology
- Structure of DNA and Gene Function
- Core Steps of Recombinant DNA Technology
- Vectors Used in Recombinant DNA Technology
- Conclusion
Introduction
1.1 Definition of Recombinant DNA Technology
Recombinant DNA (rDNA) technology is a powerful tool that has revolutionized molecular biology, genetics, and biotechnology. This technology involves combining genetic material from different sources to create new sequences that can be used for a variety of purposes, including studying gene function, producing proteins, and developing genetically modified organisms (GMOs). In zoology, rDNA technology plays a critical role in understanding animal genetics, developing vaccines, improving livestock breeds, and conserving endangered species. This essay aims to provide a comprehensive overview of recombinant DNA technology, its applications in zoology, and the ethical considerations that accompany its use.
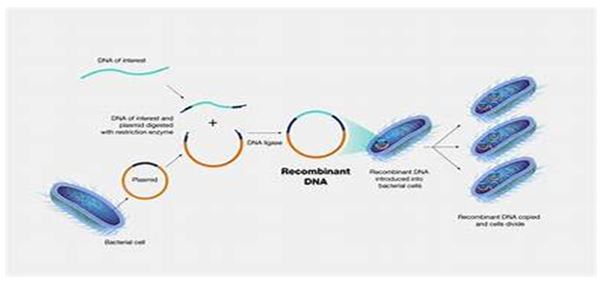
The Basics of Recombinant DNA Technology
Recombinant DNA technology is based on the principles of molecular genetics. It involves the following key steps:
1. Isolation of DNA: The first step is to extract DNA from the organism of interest. This involves breaking down cell membranes and separating the DNA from other cellular components.
2. Restriction Enzyme Digestion: The extracted DNA is then cut into fragments using restriction enzymes. These enzymes recognize specific sequences in DNA and create cuts at those sites, producing DNA fragments with “sticky” or “blunt” ends.
3. Ligation: The DNA fragment of interest is then ligated (joined) to a vector, which is a DNA molecule that can replicate independently within a host cell. Common vectors include plasmids, bacteriophages, and cosmids. This step is facilitated by the use of DNA ligase, an enzyme that catalyzes the formation of covalent bonds between the DNA fragments.
4. Transformation: The recombinant DNA is then introduced into a suitable host organism (usually bacteria, yeast, or mammalian cells) through a process known as transformation. This allows the host to express the foreign gene.
5. Screening and Selection: After transformation, it is essential to select cells that have successfully taken up the recombinant DNA. This is usually achieved using selection markers, such as antibiotic resistance genes or fluorescent proteins.
6. Expression and Analysis: The final step involves expressing the recombinant DNA and analyzing the resultant proteins or phenotypes. This can provide insights into gene function and regulation.
Applications of Recombinant DNA Technology in Zoology
Recombinant DNA technology has numerous applications within zoology, including:
1. Gene Cloning and Gene Function Studies: Gene cloning allows researchers to create copies of specific genes, enabling in-depth studies on their functions. This can help uncover genetic causes of diseases, study gene regulation, and explore evolutionary relationships.
2. Transgenic Animals: One of the most significant applications of rDNA technology is the creation of transgenic animals. These animals have had foreign DNA introduced into their genomes, allowing researchers to study gene function and expression in a whole-organism context. For example, the creation of transgenic mice has facilitated research into cancer, neurodegenerative diseases, and other complex biological processes.
3. Production of Biological Products: rDNA technology is used to produce proteins, hormones, and antibodies for therapeutic purposes. For instance, insulin used by diabetics is often produced using recombinant bacteria. Additionally, various monoclonal antibodies used in diagnostics and therapeutics are also created using these techniques.
4.Improving Livestock: In agriculture, recombinant DNA technology is utilized to enhance desirable traits in livestock, such as growth rates, disease resistance, and reproductive performance. This can lead to increased productivity and sustainability in animal farming. For example, genetically modified salmon with enhanced growth hormones reach market size more rapidly than their non-modified counterparts.
5. Vaccination and Disease Resistance: rDNA technology has also transformed vaccine development. By inserting genes encoding antigens from pathogens into vectors, researchers can produce vaccines that elicit strong immune responses. For example, recombinant vaccines for diseases like foot-and-mouth disease in livestock have improved animal health and productivity.
6. Conservation of Endangered Species: Genetic tools have been developed to aid in the conservation of endangered species. Techniques such as cloning and genetic rescue help maintain genetic diversity and increase the population size of endangered species. For example, efforts have been made to clone endangered species using rDNA technology, as seen in attempts to clone the black-footed ferret.
7. Gene Editing: Technologies such as CRISPR-Cas9 allow for targeted gene editing, enabling researchers to make precise changes to the genome. This has vast implications for gene function studies and the creation of genetically modified organisms (GMOs). Potential applications include the elimination of genetic diseases in animals and enhancement of traits such as disease resistance.
Ethical Considerations
While rDNA technology has immense potential, it also raises ethical questions and societal concerns. Some of the major considerations include:
1.Animal Welfare: The creation of transgenic animals and other genetic modifications can raise questions about animal welfare. Concerns include the potential for unintended consequences, such as health issues, altered behavior, and suffering associated with genetic changes.
2.Biodiversity and Ecosystem Impact: The introduction of genetically modified organisms into the environment raises concerns about their potential impact on ecosystems and biodiversity. For example, if genetically modified animals escape into the wild, they could disrupt local populations and ecosystems.
3. Food Safety and Public Health: The use of transgenic animals in food production raises questions about food safety, allergenicity, and long-term health effects. Regulatory frameworks and rigorous testing are necessary to ensure the safety of genetically modified food products.
4. Ethical Treatment of Animals: The ethical implications of using animals for genetic research are significant. Researchers must consider the humane treatment of experimental animals and use alternatives whenever possible.
5.Public Perception and Acceptance: Public perception of rDNA technology varies widely. While some individuals embrace the benefits of genetic engineering, others express concerns about safety, ethics, and the potential for misuse. Effective communication and transparency are crucial for fostering public trust.
1.2 Historical Context
The origins of recombinant DNA technology can be traced back to the early 1970s when researchers, including Paul Berg, Herbert Boyer, and Stanley Cohen, pioneered techniques for cutting and splicing DNA. The groundbreaking experiments involved the use of restriction enzymes, proteins that act as molecular scissors to cut DNA at specific sequences. By combining DNA from different organisms, these scientists demonstrated that it was possible to create new genetic combinations. Their work laid the foundation for a plethora of applications and ignited discussions about the ethical implications of genetic manipulation.
2.0 Key Concepts in Recombinant DNA Technology
Recombinant DNA (rDNA) technology encompasses various fundamental principles and techniques that allow scientists to manipulate genetic material. Understanding these key concepts is essential for grasping how rDNA technology operates and its applications. Here are the primary concepts involved:
1. Molecular Genetics Overview
Molecular genetics is the field of biology that uses the principles of genetics and molecular biology to study gene structure and function. It integrates both genetic and biochemical methods to investigate how genes are expressed and regulated.
2. Basic Terminology
DNA (Deoxyribonucleic Acid): The molecule that carries genetic information in living organisms, consisting of two strands that form a double helix.
Gene: A specific segment of DNA that encodes for a particular protein or functional RNA.
Genome: The complete set of genetic material in an organism.
Plasmid: A small, circular piece of DNA that can replicate independently from chromosomal DNA, often used as a vector in genetic engineering.
3. Structure of DNA
DNA is composed of nucleotides, each containing a phosphate group, a sugar (deoxyribose), and a nitrogenous base (adenine, thymine, cytosine, or guanine). The sequence of these bases encodes genetic information. The double helix structure of DNA comprises two anti-parallel strands, held together by hydrogen bonds between complementary base pairs (A-T and C-G).
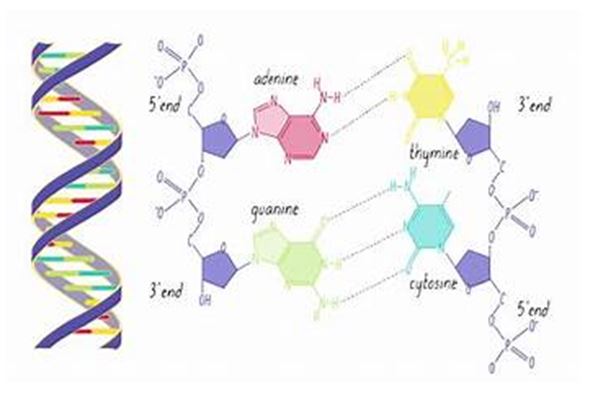
4. Gene Function
Transcription: The process by which a gene’s DNA sequence is copied into messenger RNA (mRNA), which then exits the nucleus into the cytoplasm.
Translation: The process where ribosomes synthesize proteins using the sequence of the mRNA, translating the genetic code into an amino acid sequence.
Gene Regulation: Mechanisms that control the expression of genes, including transcription factors, enhancers, and repressors.
5. Vectors
Vectors are DNA molecules used to deliver genetic material into cells. They can carry foreign DNA into host cells, allowing for replication and expression.
Types of Vectors include plasmids, bacteriophages, cosmids, and artificial chromosomes.
6. Restriction Enzymes
Restriction enzymes (also known as restriction endonucleases) are proteins that cut DNA at specific sequences, producing fragments with “sticky” or “blunt” ends. This allows for precise manipulation of DNA and is crucial for inserting genes into vectors.
7. Ligation
Ligation is the process of joining together DNA fragments. DNA ligase, an enzyme, is used to form covalent bonds between the sugar-phosphate backbone of DNA strands, effectively attaching the foreign DNA to the vector.
8. Transformation
Transformation is the process by which a host cell takes up the recombinant DNA. This can be achieved using various methods, including heat shock, electroporation, and viral infection, depending on the type of host organism.
9. Screening and Selection
After transformation, it is crucial to identify and select the cells that have successfully incorporated the recombinant DNA. Common methods for screening include antibiotic selection (where only transformed cells survive in the presence of an antibiotic) and marker genes (such as fluorescent proteins).
10. Cloning
Cloning refers to the process of creating copies of DNA fragments or whole organisms. In the context of rDNA technology, it often pertains to replicating a specific gene for further study or use.
11. Gene Editing
Gene editing technologies, such as CRISPR-Cas9, allow for precise modifications of the genome. These advancements enable targeted alterations, corrections, or deletions of specific DNA sequences, leading to potential therapeutic applications.
12. Applications in Research and Medicine
rDNA technology has applications in various fields, including agriculture (genetically modified crops), medicine (production of insulin and vaccines), and conservation (cloning and genetic rescue programs for endangered species).
3.0 Core Steps of Recombinant DNA Technology
Recombinant DNA technology involves several key steps that collectively enable the manipulation of genetic material. Among these steps, DNA isolation, restriction enzyme digestion, and DNA ligation and vector construction are foundational processes. Below is a detailed overview of each step:
1. DNA Isolation
Definition:
DNA isolation, also known as DNA extraction, is the process of purifying DNA from the cells of an organism. The goal is to obtain intact DNA that can then be used in subsequent steps of recombinant DNA technology.
Process:
The isolation process typically involves several steps:
Cell Lysis: Cells are broken open using physical methods (like grinding or sonication) or chemical methods (like detergents). This step is crucial as it releases the DNA into solution alongside other cellular components such as proteins, lipids, and RNA.
Removal of Proteins and Contaminants: Enzymes such as proteases can be added to digest proteins. Organic solvents, such as phenol-chloroform, or precipitation with alcohol (such as isopropanol or ethanol) are often used to separate the DNA from proteins and other debris. The DNA can be precipitated out of solution for further purification.
Purification and Quantification: The isolated DNA is then washed and resuspended in a suitable buffer for downstream applications. Quantification can be performed using spectrophotometric methods to determine DNA concentration and purity.
Importance:
This step is crucial as the quality of DNA extracted directly influences the success of subsequent experiments. Contaminated or degraded DNA may lead to inefficient cloning or unreliable results in applications such as gene expression analysis
2. Restriction Enzyme Digestion
Definition:
Restriction enzyme digestion involves the cutting of DNA at specific sequences by restriction enzymes (also known as restriction endonucleases). This allows researchers to create DNA fragments that can be manipulated for cloning purposes.
Process:
Selection of Restriction Enzymes: Each restriction enzyme recognizes specific DNA sequences (usually palindromic) and creates cuts at those sites. Researchers select enzymes based on the desired fragment size and the presence of specific recognition sites in the target DNA.
Digestion Reaction: The isolated DNA is mixed with the chosen restriction enzymes, along with a suitable buffer that provides the optimal conditions (like pH and salt concentration) for the enzyme’s activity. The mixture is incubated at a specific temperature for a certain amount of time to enable the enzymes to cut the DNA.
Resulting Fragments: The digestion produces DNA fragments with either “sticky” or “blunt” ends, depending on the type of cut made by the restriction enzyme. Sticky ends are preferable for cloning because they facilitate the ligation of DNA fragments
Importance:
This step allows for the precise manipulation of DNA, enabling the isolation of specific genes or sequences of interest. The creation of compatible ends enhances the effectiveness of subsequent ligation steps, ensuring successful incorporation of foreign DNA into vectors.
3. DNA Ligation and Vector Construction
Definition:
DNA ligation involves joining two pieces of DNA together, typically in the context of DNA cloning where the goal is to insert a fragment of interest into a vector—a DNA molecule that will carry the fragment into a host cell.
Process:
Choice of Vector: Vectors can be plasmids, bacteriophages, or cosmids, which are selected based on the size of the DNA fragment to be cloned, the type of host cells, and the desired applications (like expression, replication, or stability).
Preparation of Vector: The vector must be linearized by cutting it with the same restriction enzyme used to digest the target DNA. This ensures that the ends of the vector and the insert fragment will be compatible.
Ligation Reaction: A DNA ligase enzyme is used to catalyze the formation of covalent phosphodiester bonds between the sugar-phosphate backbones of the insert and vector DNA, effectively joining them together. This reaction commonly occurs at room temperature for a specified duration.
Resulting Construct: The resulting DNA molecule, now a recombinant vector containing the foreign DNA, is constructed for the next steps (e.g., transformation into a host organism).
Importance:
The ligation step is critical because successful joining of the insert with the vector is essential for the propagation of the inserted gene in host cells. A well-constructed recombinant vector allows for the effective expression, analysis, or further manipulation of the introduced gene.
4.0 Vectors Used in Recombinant DNA Technology
Recombinant DNA (rDNA) technology is a cornerstone of molecular biology and biotechnology, allowing scientists to manipulate genetic materials from various organisms. A crucial component of this technology is the use of vectors—DNA molecules that facilitate the introduction of foreign genetic material into host cells. Vectors serve as vehicles to transfer, replicate, and express the inserted genes, playing a vital role in applications ranging from gene cloning to the development of genetically modified organisms (GMOs) and therapeutic proteins.
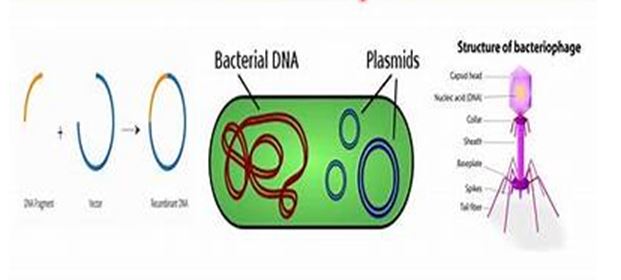
This article will delve into the types of vectors used in recombinant DNA technology, their functions, and their specific applications within the broader context of life sciences. By understanding the various vector systems, scientists can select the most appropriate vector for their research or industrial needs.
1. Overview of Vectors
Vectors are DNA molecules that can carry foreign DNA fragments. They are essential in gene cloning, as they help researchers insert new genes into host cells, where these genes can be expressed. Vectors are designed to facilitate replication and expression of the foreign DNA and often contain specific features that allow for their identification and manipulation within the host.
Essential Characteristics of Vectors
For a DNA molecule to function effectively as a vector, it should possess certain features:
Origin of Replication (ori): This region is crucial for the vector’s ability to replicate independently within the host cell. It ensures that the vector and the attached plasmid DNA are copied during cell division.
Selectable Marker Genes: These genes confer resistance to antibiotics or other selection agents, enabling researchers to identify which host cells have successfully taken up the vector. Common markers include genes for antibiotic resistance (e.g., ampicillin, tetracycline).
Multiple Cloning Sites (MCS): An MCS is a short region containing several restriction enzyme recognition sites, allowing for the insertion of foreign DNA. It facilitates the directional cloning of the gene of interest.
Promoter Region: This sequence drives the expression of the inserted gene. Depending on the vector’s purpose, it may be constitutive (always active) or inducible (activated under specific conditions).
Expression Control Elements: In expression vectors, elements such as enhancers and terminators are included to regulate the efficient transcription and translation of the cloned gene.
2. Types of Vectors
Several types of vectors are utilized in recombinant DNA technology, each designed for specific applications. Here, we will explore the most common vector types:
2.1 Plasmid Vectors
Plasmids are circular, double-stranded DNA molecules naturally found in bacteria. They replicate independently of chromosomal DNA and are among the most widely used vectors in rDNA technology.
Characteristics of Plasmid Vectors:
Small Size: Plasmids can be relatively small, making them easy to manipulate.
Autonomous Replication: They replicate independently within bacterial cells due to their origin of replication.
Selectable Markers: They often carry genes that confer antibiotic resistance.
Applications:
Gene Cloning: Plasmids are commonly used to clone fragments of DNA for analysis or expression in bacteria.
Protein Production: Plasmids can be designed to express proteins in bacterial systems, facilitating the study of proteins or production of pharmaceutical compounds.
Example: pBR32
One of the first widely used plasmid vectors, pBR322 contains genes for ampicillin and tetracycline resistance and multiple restriction sites.
2.2 Bacteriophage Vectors (Lambda Vectors)
Bacteriophages (viruses that infect bacteria) can also serve as vectors. Lambda phage, in particular, has been extensively used for cloning large DNA fragments.
Characteristics of Bacteriophage Vectors:
Capacity for Large Inserts: Lambda vectors can incorporate larger DNA fragments compared to plasmids, allowing for the cloning of entire genes or operons.
Life Cycle: Bacteriophages can undergo lytic or lysogenic cycles, allowing for various applications in molecular genetics.
Applications:
Construction of Genomic Libraries: Lambda vectors are commonly used in the creation of libraries that include a range of DNA sequences from a particular organism.
Example: Lambda ZAP
A lambda vector designed for the cloning of cDNA libraries. It allows for both cloning and expression of the genes carried within the vector.
2.3 Cosmids
Cosmids are hybrids between plasmids and bacteriophage lambda. They combine features of both, allowing for the cloning of larger DNA fragments than traditional plasmids.
Characteristics of Cosmids:
Large Insert Size: Cosmids can accommodate DNA inserts ranging from 35 to 45 kilobases, enabling the cloning of larger genes or genomic regions.
Transduction Ability: Using phage features, cosmids can be introduced into host cells via transduction.
Applications:
Genomic Library Construction: They are useful for constructing genomic libraries of larger organisms and are crucial in sequencing projects.
2.4 Artificial Chromosomes
Artificial chromosomes can be generated from either yeast (YACs) or bacteria (BACs). These vectors can carry large inserts of genetic material, facilitating the genomic mapping and sequencing projects.
Characteristics of Artificial Chromosomes:
Capacity for Large Inserts: BACs can carry 100-300 kb of DNA, while YACs can accommodate even larger fragments, up to 1 megabase.
Eucaryotic Constructs Available: YACs replicate in yeast, allowing for the study of eukaryotic genes in a eukaryotic system.
Applications:
Genome Mapping and Sequencing: Artificial chromosomes are critical for genomic sequencing efforts, providing a means to clone and analyze large segments of DNA.
Studying Eukaryotic Genes: They can also be used for functional studies involving eukaryotic genes and regulatory elements.
Examples:
Bacterial Artificial Chromosomes (BACs): Derived from F plasmids; used extensively in the Human Genome Project for cloning large genomic DNA segments.
Yeast Artificial Chromosomes (YACs): Initially developed to clone large human genomic fragments.
2.5 Expression Vectors
Expression vectors are specially designed to express a cloned gene within a host cell, producing the corresponding protein. They incorporate all necessary elements for transcription and translation.
Characteristics of Expression Vectors:
Promoter Sequences: They contain strong, regulated promoters that ensure high levels of gene expression.
Transcriptional and Translational Signals: Include sequences for ribosome binding, transcription termination, and other regulatory elements.
Applications:
Protein Production: Widely used in biotechnology for the production of recombinant proteins, including hormones, enzymes, and antibodies.
Example: pET System
A popular expression system that uses T7 bacteriophage promoters to drive expression in E. coli. It is widely utilized for producing high quantities of recombinant proteins.
3. Vector Selection Criteria
When choosing a vector for a specific application, researchers must consider several factors:
1. Insert Size: The ability of the vector to accommodate the desired size of the foreign DNA is paramount. For smaller inserts, plasmids are often sufficient; for larger fragments, cosmids or artificial chromosomes may be necessary.
2. Host Organism: The choice of host organism can influence the selection of a vector. For instance, bacterial vectors are optimized for E. coli, while yeast vectors are used for yeast host systems.
3. Purpose of Cloning: The ultimate goal of the cloning experiment (e.g., gene expression, protein production, sequencing) will dictate the vector design, including the need for specific regulatory elements.
4. Ease of Use: Factors such as cloning efficiency, cost, and available protocols for handling the vector may also influence the choice of vector.
4. Applications of Vectors in Different Fields
4.1 Biotechnology
Biotechnology often utilizes vectors to create recombinant proteins, such as insulin and growth factors, which are crucial for various medical treatments. Expression vectors are key to producing these proteins in suitable host systems.
4.2 Genetic Engineering
Vectors are essential in genetic engineering, including the development of GMOs. By incorporating genes that confer beneficial traits, such as pest resistance, vectors are instrumental in agricultural biotechnology.
4.3 Gene Therapy
In gene therapy research, vectors are used to deliver therapeutic genes to target cells. Commonly, viral vectors (derived from adenoviruses or lentiviruses) are employed for efficient gene delivery in human cells.
4.4 Research and Development
Vectors are crucial tools in basic research for cloning genes, creating gene knockout models, and performing functional studies on gene expression. Researchers employ a variety of vectors to explore gene functions and interactions systematically.
5. Challenges and Considerations
Despite the advantages of using vectors in recombinant DNA technology, researchers face several challenges:
1.Insertional Mutagenesis: Random insertion of vectors may disrupt essential genes or regulatory elements, leading to unintended consequences. This is particularly important in applications such as gene therapy.
2. Host Compatibility: Not all vectors are compatible with all host organisms. Researchers must select vectors and host systems that work well together to ensure successful expression and function.
3. Stability of Inserts: Long-term stability of the inserted DNA can be a concern, particularly in plasmids. Some constructs may undergo rearrangements or deletions over time.
4. Regulatory Barriers: The use of vectors for creating GMOs or gene therapies is subject to regulatory scrutiny, requiring extensive safety assessments and compliance with ethical standards.
6. Future Directions
The field of recombinant DNA technology continues to evolve, with advancements in vector design and applications. Future directions may include:
1.Novel Vector Development: Creating vectors with enhanced capabilities, such as improved stability, expression efficiency, and versatility for different host systems.
2. Viral Vectors: Researchers are increasingly developing safer and more efficient viral vectors for gene therapy, focusing on reducing immune responses while maintaining delivery efficacy.
3. CRISPR-Compatible Vectors: As CRISPR technology advances, the development of vectors that can facilitate gene editing in a precise and targeted manner will further enhance the possibilities for genetic manipulation.
4.Synthetic Biology Applications: The integration of synthetic biology concepts into vector design may lead to innovative applications, such as creating custom-designed organisms with specific functions or characteristics.
5.0 Techniques Associated with Recombinant DNA Technology
Recombinant DNA technology has revolutionized molecular biology and genetics, enabling scientists to isolate, manipulate, and analyze genes in ways that were previously unimaginable. This essay will provide an overview of the various techniques that form the foundation of recombinant DNA technology, detailing the principles, processes, and applications of each method. The techniques discussed will include DNA isolation, restriction enzyme digestion, ligation, transformation, screening and selection, polymerase chain reaction (PCR), DNA sequencing, and gene editing technologies such as CRISPR-Cas9.
1. DNA Isolation
Principle: The first step in recombinant DNA technology involves isolating DNA from the organism of interest. This process must be performed gently to ensure the integrity of the DNA, as it can be easily degraded by nucleases.
Process:
Cell Lysis: Cells are broken open (lysed) using detergents or enzymes. This step releases the contents of the cells, including the DNA, into the solution.
Removal of Proteins and Contaminants: Proteins and other cellular debris are removed through various methods, such as centrifugation or the use of proteolytic enzymes (e.g., proteinase K).
Precipitation of DNA: The DNA is precipitated out of the solution using alcohol (ethanol or isopropanol). This process involves mixing the aqueous solution containing the DNA with alcohol, which causes the DNA to form a visible, solid mass.
Purification: The DNA can be further purified to remove any remaining contaminants, ensuring it is suitable for subsequent experiments.
Applications: Isolated DNA can be used for cloning, sequencing, and various forms of genetic analysis.
2. Restriction Enzyme Digestion
Principle: Restriction enzymes (or restriction endonucleases) are proteins that cleave DNA at specific nucleotide sequences. Each enzyme recognizes a unique sequence and makes cuts at or near that site, which is critical for creating fragments that can be manipulated in subsequent steps.
Process:
Selection of Restriction Enzymes: Appropriate restriction enzymes are selected based on the target DNA sequence. Commonly used enzymes include EcoRI, HindIII, and BamHI.
Digestion Reaction: The isolated DNA is mixed with the chosen restriction enzyme and a buffer solution. The reaction conditions (temperature, time, and ionic strength) are optimized according to the enzyme’s requirements.
Fragment Analysis: After digestion, the resulting DNA fragments can be analyzed through techniques such as gel electrophoresis, which separates the fragments based on size, allowing verification of the digestion process.
3. Ligation
Principle: Ligation involves joining two DNA fragments together. This step is crucial for creating recombinant DNA molecules that can be introduced into host cells.
Process:
Preparation of DNA Fragments: The DNA fragments that need to be joined—often one containing the gene of interest and the other a vector—must be digested with the same or compatible restriction enzymes to create complementary sticky or blunt ends.
Mixing with DNA Ligase: The DNA fragments are mixed with DNA ligase, an enzyme that forms covalent bonds between the phosphate backbone of the DNA strands.
Incubation: The mixture is incubated under conditions favorable for ligation, allowing the enzyme to facilitate the bond formation between the DNA fragments.
Applications: Ligation is primarily used in cloning experiments to insert genes into plasmids or other vectors, creating recombinant constructs that can be introduced into host organisms.
4. Transformation
Principle: Transformation is the process of introducing recombinant DNA into a host cell so that it can be replicated and expressed. This is typically performed in bacterial cells, commonly *Escherichia coli (E. coli), although eukaryotic cells can also be transformed.
Process:
Preparation of Competent Cells: Bacterial cells must be made competent to take up DNA. This can be achieved using chemical methods (e.g., calcium chloride treatment) or electroporation, which subjects cells to an electric field to increase permeability.
Mixing with DNA: The competent cells are mixed with the ligated DNA (recombinant DNA) and incubated on ice to promote DNA uptake.
Heat Shock or Electroporation: Cells are subjected to a heat shock (42°C for 30–60 seconds) or an electric pulse to facilitate the uptake of DNA.
Recovery: Cells are allowed to recover in a nutrient-rich medium, enabling expression of any antibiotic resistance genes carried by the vector.
5. Screening and Selection
Principle: After transformation, it is necessary to identify which cells have successfully taken up the recombinant DNA. Screening and selection techniques are employed to distinguish transformed cells from those that did not incorporate the DNA.
Process:
Use of Selection Markers: Most cloning vectors contain antibiotic resistance genes that allow for the selection of successful transformants. When grown on agar plates containing the corresponding antibiotic, only transformed cells will survive.
Colony Screening: Colonies that grow on selective media can be further screened using methods such as:
Colony PCR: To amplify the inserted gene from bacterial colonies for confirmation.
Restriction Digestion: Isolate plasmid DNA from successful colonies and digest with restriction enzymes to check for the presence and size of the insert.
Sequencing: Confirm the correct sequence of the inserted DNA by sequencing the plasmid DNA.
Applications: Screening and selection are crucial for identifying successful clones in gene cloning experiments and ensuring that the desired genetic material is present.
6. Polymerase Chain Reaction (PCR)
Principle: PCR is a technique developed to amplify specific DNA sequences exponentially, allowing researchers to obtain sufficient quantities of DNA for further analysis. This technique employs cycles of denaturation, annealing, and extension.
Process:
Denaturation: The double-stranded DNA template is heated to a high temperature (typically 94–98°C), causing the strands to separate.
Annealing: The temperature is lowered (typically 50–65°C) to allow specific primers to bind (anneal) to their complementary sequences on the target DNA.
Extension: The temperature is raised again (usually to around 72°C) for Taq polymerase (a heat-stable DNA polymerase) to synthesize new DNA strands by extending from the primers.
Cycling: This process is repeated for 25–40 cycles, resulting in an exponential increase in the amount of target DNA.
Applications: PCR is used for various applications such as cloning, gene expression analysis, detection of pathogens, and genetic fingerprinting.
7. DNA Sequencing
Principle: DNA sequencing determines the precise order of nucleotides in a DNA molecule. It is essential for confirming the identity of cloned genes and for applications in genomics and molecular biology.
Process:
Sanger Sequencing: The traditional method using dideoxynucleotides (ddNTPs) that terminate DNA synthesis. Each reaction produces fragments of different lengths that correspond to the nucleotides in the template DNA.
Next-Generation Sequencing (NGS): High-throughput technologies that allow massive parallel sequencing, enabling the sequencing of entire genomes rapidly and cost-effectively.
Bioinformatics Analysis: The generated sequences are analyzed using bioinformatics tools to identify genes, variants, and other important genomic features.
6.0 Conclusion
Recombinant DNA technology has fundamentally transformed the fields of molecular biology, biotechnology, and genetics, enabling unprecedented advancements in our understanding of life at a molecular level. By allowing scientists to manipulate genetic material, this technology has opened doors for groundbreaking applications, ranging from creating transgenic animals that serve as models for human diseases to producing therapeutic proteins and vaccines that have improved health outcomes worldwide. The ability to clone genes, edit genomes, and develop genetically modified organisms has significant implications for agriculture, medicine, and conservation.
However, the incorporation of recombinant DNA technology into practical applications also raises critical ethical and societal concerns. Issues related to animal welfare, environmental impacts, food safety, and public perception must be addressed thoughtfully and transparently. As research and applications continue to evolve, it is essential for stakeholders—including scientists, policymakers, and the public—to engage in open dialogues about these challenges.
The future of recombinant DNA technology holds immense promise, with emerging techniques such as CRISPR-Cas9 and advancements in synthetic biology poised to further enhance our capabilities. Balancing innovation with ethical stewardship will be crucial in ensuring that these genetic tools are used responsibly. By prioritizing ethical considerations and ensuring public engagement, we can harness the full potential of recombinant DNA technology to improve health, agriculture, and environmental sustainability while fostering a society that supports science and research. Looking ahead, the ongoing exploration and application of this technology will shape the future of biology, agriculture, and medicine, paving the way for solutions to some of the most pressing challenges we face today.
Click for further information
https://a-zstudentpdfprojectfilesolutions.com/category/zoology-knowledge/
